> library(lattice)
Warning message:
程辑包‘lattice’是用R版本3.4.3 来建造的
> library(caret)
载入需要的程辑包:ggplot2
Warning messages:
1: 程辑包‘caret’是用R版本3.4.3 来建造的
2: 程辑包‘ggplot2’是用R版本3.4.3 来建造的
> library(dplyr)
载入程辑包:‘dplyr’
The following objects are masked from ‘package:stats’:
filter, lag
The following objects are masked from ‘package:base’:
intersect, setdiff, setequal, union
Warning message:
程辑包‘dplyr’是用R版本3.4.3 来建造的
> library(elasticnet)
载入需要的程辑包:lars
Loaded lars 1.2
> library(lars)
> sim.dat=read.csv("https://raw.githubusercontent.com/happyrabbit/DataScientistR/master/Data/SegData.csv")
>
> ymad=mad(na.omit(sim.dat$income))
>
> zs=(sim.dat$income-mean(na.omit(sim.dat$income)))/ymad
>
> idex=c(which(na.omit(zs>3.5)),which(is.na(zs)))
>
> sim.dat=sim.dat[-idex,]
>
> xtrain=dplyr::select(sim.dat,Q1:Q10)
>
> ytrain=sim.dat$income
>
> set.seed(100)
>
> ctr1=trainControl(method="cv",number=10)
>
> plsTune=train(xtrain,ytrain,method="pls",tuneGrid=expand.grid(.ncomp=1:10))
>
> plsTune=train(xtrain,ytrain,method="pls",tuneGrid=expand.grid(.ncomp=1:10),trControl=ctr1)
>
> pluTune
Error: object 'pluTune' not found
> plsTune
Partial Least Squares
772 samples
10 predictor
No pre-processing
Resampling: Cross-Validated (10 fold)
Summary of sample sizes: 694, 696, 696, 696, 694, 695, ...
Resampling results across tuning parameters:
ncomp RMSE Rsquared MAE
1 28106.40 0.6553646 19957.91
2 24852.89 0.7385908 16142.37
3 23594.19 0.7679501 14507.19
4 23442.31 0.7713064 13940.47
5 23407.49 0.7721321 13848.48
6 23409.49 0.7720994 13838.38
7 23408.15 0.7721470 13835.54
8 23408.56 0.7721433 13835.52
9 23408.46 0.7721447 13835.46
10 23408.46 0.7721448 13835.48
RMSE was used to select the optimal model using the smallest value.
The final value used for the model was ncomp = 5.
> plot(plsImp,top=10,scales=list(y=list(cex=.95)))
可以看到,Q1Q2,Q3的重要性明显高于其他变量
下面我们对主成分回归进行调优,调优参数依旧是模型中成分的数目
> set.seed(100)
> pcrTune=train(x=xtrain,y=ytrain,method="pcr",tuneGrid=expand.grid(.ncomp=1:10),trControl=ctr1)
> pcrTune
Principal Component Analysis
772 samples
10 predictor
No pre-processing
Resampling: Cross-Validated (10 fold)
Summary of sample sizes: 696, 693, 694, 694, 696, 695, ...
Resampling results across tuning parameters:
ncomp RMSE Rsquared MAE
1 45957.59 0.03243035 36598.90
2 32459.79 0.52199913 24040.87
3 23235.11 0.75774323 14515.67
4 23262.34 0.75735221 14545.16
5 23152.29 0.75956717 14231.64
6 23133.20 0.76003827 14130.41
7 23113.93 0.76048652 14128.53
8 23114.64 0.76045268 14129.77
9 22991.03 0.76283052 13801.16
10 22976.35 0.76308486 13726.14
RMSE was used to select the optimal model using the smallest value.
The final value used for the model was ncomp = 10.
> plsResamples=plsTune$results
> plsResamples$Model="PLS"
> pcrResamples=pcrTune$results
> pcrResamples$Model="PCR"
> plsPlotData=rbind(plsResamples,pcrResamples)
> xyplot(RMSE~ncomp,data=plsPlotData,xlab="# Components",ylab="RMSE (Cross-Validation)",auto.key=list(columns=2),groups=Model,type=c("o","g"))
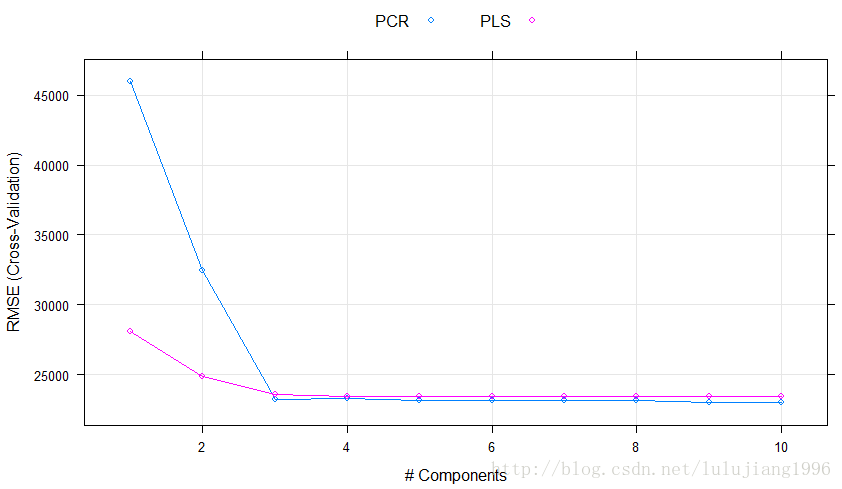
如上,图形表明:用前3个成分给出的模型效果和用更多成分给出的模型效果依旧没有太大差别了。